Final Technical Report - Molecular profile analysis of vaccines

Preface
First of all we would like to thank the very useful comments provided by those who have reviewed the results of the analyses carried out in the context of scientific research activities relating to Priorix tetra and Infanrix Hexa products. The critical issues presented were indeed very useful in order to add technical-scientific integrations able to clarify the work done. We believe that only through a healthy communion of scientific visions can conclusions be reached on the data obtained that can be useful for the entire scientific community and for the people who address it.
1. State of the art
Preliminary studies (screening not subject to confirmation) of biomolecular, metabolomic and proteomic profile, performed on Priorix Tetra and Infanrix Hexa products have led to a composition framework summarized in the following points:
- Presence of different analytical signals that can not be associated with known compounds through research on the Metlin 1-2 and KEGG3 databases. Therefore, a picture emerged associated with a considerable complexity in the composition of commercial products.
- Presence of proteins not declared in a leaflet in the Priorix Tetra product. The latter can potentially be associated with residues of the production process
- Non-detection of the antigens declared within the Infanrix Hexa product. The analysis technique consisted of enzymatic digestion with trypsin associated with mass spectrometry techniques. 4-5
These data have raised several comments especially regarding point C - Protein detection is in fact carried out using a standard approach, internationally recognized for over 10 years 4, of digestion through the trypsin enzyme 4. The peptides thus obtained are chromatographically separated and analyzed by mass spectrometry 4-5. The main observation has been inherent in the fact that in the vaccines there are aluminum-based adjuvants that could potentially inhibit the enzymatic digestion process. The data acquired subsequently made it possible to provide substantial clarifications, especially with regard to the complaint expressed in point C.
2. New insights and analysis
2.1 Insights related to the analysis of the Infanrix Hexa product
Before proceeding with the illustration of the new acquired data concerning Hexyon and Gardasil 9 vaccines, it is essential to answer the question concerning the doubt raised regarding the inhibition of the proteolytic activity of trypsin caused by the presence of aluminum-based adjuvants in the Infanrix Hexa vaccine. In this regard it must be specified that a digestion control is always present within the tryptic digestion. In fact, the trypsin used to perform digestion, although engineered to prevent autolysis, has a small percentage of the latter that in the case of enzymatic activity leads to the obtainment of the fragment having m/z 842 and the following peptide sequence : VATVSLPR. Said fragment was actually detected within the tryptic digestion of the Infanrix Hexa product as verifiable by the ion extraction chromatogram (Figure 1).
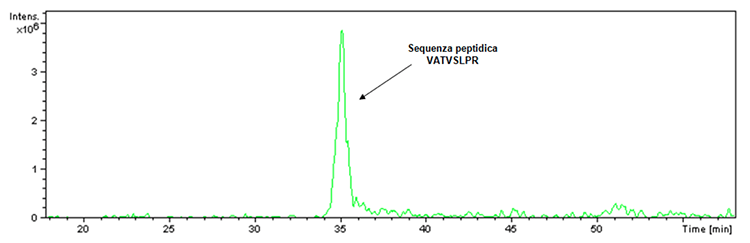
Figure 1: Ion-associated ionic extraction chromatogram with m/z 842 ratio found in the sample of the Infanrix Hexa product lot (batch No. A21CD072D).
Furthermore, an external control is performed by digesting hemoglobin, in order to further verify the goodness of the lot of trypsin used. Hemoglobin, analyzed in the analysis section in which the product was monitored, was recognized with a significant statistical score (loge < - 100). These data confirmed the fact that the enzyme activity was present.
2.2 New analyses concerning Hexyon and Gardasil products
The analysis of Hexyon and Gardasil 9 products led to the detection of complex molecular profiles. In this case, however, the presence of most of the antigens reported in the information leaflet was detected. They were detected by tryptic digestion and in the presence of adjuvants.
This fact further strengthens the evidence that the triptica digestion reaction is not inhibited in the presence of adjuvants. In the case of Hexyon and Gardasil 9 vaccines, the complexity of molecular profile was mainly attributed to the presence of numerous species, with low molecular weight, not identifiable by the reference databases Metlin 1-2- and KEGG 3.
3. Conclusions and final considerations
The analyses carried out led to the conclusion of the following:
- The molecular profile of the vaccines analyzed is generally complex and largely unknown.
- There are protein contaminations, not declared in the leaflet, whose composition is variable.
- In several cases the antigens declared in the leaflet have not been detected. This fact could be attributed to several factors. Among the latter, we can consider the sensitivity of the method used. However, we feel we can exclude the phenomenon of digestion inhibition due to the presence of adjuvants in the formulation of the vaccine. In fact, the enzymatic activity is mainly confirmed by the presence of fragments of tryptic autolysis, within the solutions of the digested vaccines (internal control).
4. Future studies
Further studies will be carried out as part of the research and development activities aimed at investigating the following aspects:
- macromolecular composition associated with solid residues present in vaccines (MALDI TOF MS analysis); 5
- evaluation of the concentration of the metals present in the products.
- Second level analysis to confirm the presence of toxic compounds detected in the screening phase. Their concentration will then be related to the toxicity of the same according to what stated on the international safety data sheets. The second level analyzes will be carried out in compliance with the European directive EU 2002/657/EC, useful to guarantee high quality standards in the mass spectrometry sector. 6
Loretta Bolgan
5. Bibliographic References
- Autenhahn R, Cho K, Uritboonthai W, Zhu Z, Patti G, Siuzdak G (September 2012). "An accelerated workflow for untargeted metabolomics using the METLIN database". Nature Biotechnology. 30: 826–828. doi:10.1038/nbt.2348. PMC 3666346. PMID 22965049.
- Smith CA, I'Maille G, Want EJ, Qin C, Trauger SA, Brandon TR, Custodio DE, Abagyan R, Siuzdak G (December 2005). "METLIN: a metabolite mass spectral database" (PDF). Ther Drug Monit. 27 (6): 747–51. doi:10.1097/01.ftd.0000179845.53213.39. PMID 16404815.
- Kanehisa M (2013). "Chemical and genomic evolution of enzyme-catalyzed reaction networks". FEBS Lett. 587 (17): 2731–7.
- Cristoni S, Bernardi LR. "Bioinformatics in mass spectrometry data analysis for proteomics studies." Expert Rev Proteomics. 2004 Dec;1(4):469-83.
- Cristoni S, Bernardi LR. "Development of new methodologies for the mass spectrometry study of bioorganic macromolecules." Mass Spectrom Rev. 2003 Nov-Dec;22(6):369-406.
- Cristoni S, Dusi G, Brambilla P, Albini A, Conti M, Brambilla M, Bruno A, Di Gaudio F, Ferlin L, Tazzari V, Mengozzi S, Barera S, Sialer C, Trenti T, Cantu M, Rossi Bernardi L, Noonan DM. " SANIST: optimization of a technology for compound identification based on the European Union directive with applications in forensic, pharmaceutical and food analyses." J. Mass Spectrom. 2017 Jan;52(1):16-21. doi: 10.1002/jms.3895.
Download: CORVELVA-Final-Technical-Report.pdf
Translated by team CLiVa - www.clivatoscana.com

